Wastewater treatment plant operators have relied on a variety of monitoring tools — DO, pH, SV30/SVI, sludge depth — to keep facilities running efficiently and within permit requirements. While microscopic exams and ATP measurements provide insight into the health of the biological unit’s biomass, these techniques provide limited insight into the actual bacteria required for efficient waste treatment.
Recent advances in molecular biology now make it possible and affordable to understand and track the key bacteria — both good and bad — that impact wastewater treatment plant performance. Two of these advances, microbial community analysis (MCA) and quantitative polymerase chain reaction (qPCR) — which is the molecular biology equivalent of a plate count on highly selective agars — give operators critical insight into the composition and health of their mixed liquor suspended solids or biomass, particularly during upsets or when introducing operational changes or new influents.
Microbial community analysis
An MCA is in effect a total microbial census. A biomass sample is used for high-throughput, next-generation sequencing of a genetic segment present in all bacteria. By looking at the variations in the genetic sequences, bioinformatics software identifies and counts the types of bacteria present in a sample. The results are further analyzed to identify the relative population sizes of important groups, such as:
● Ammonia oxidizers & nitrite oxidizers (AOB & NOB)
● Denitrifiers
● Phosphorus accumulators (PAO)
● Odor-causing sulfate reducers (SRB)
● Filamentous, bulking or foaming bacteria
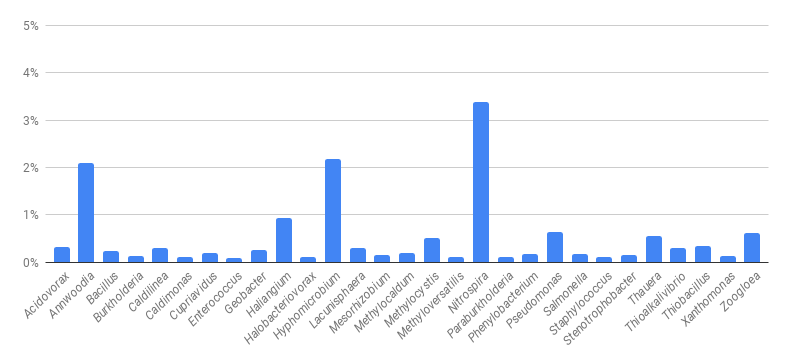
The above figure is an example of an MCA for a “healthy” wastewater treatment plant, showing bacteria present at 0.1% or higher. Nitrifiers, primarily represented by Nitrospira, and denitrifiers (Hyphomicrobium, Pseudomonas and Thauera) are abundant. Significant levels of Annwoodia and Thiobacillus, both sulfur-oxidizing bacteria, indicate that the influent contains significant levels of organosulfur and sulfide compounds. And depending on plant conditions and the exact strain, Caldilinea can appear as either Type 0914 or Type 0803 filaments under a microscope.
The microbial community analysis can help operators implement critical decisions. For example, a wastewater treatment plant at a chicken processing facility was losing ammonia removal during winter. MCA revealed that the facility could maintain a low level of nitrifiers (around 0.4% to 0.6% of the biomass) by trucking in nitrifying sludge from neighboring facilities. However, this nitrifier population was unable to sufficiently remove ammonia to meet permit. As the nitrifiers were present, this suggested that slow growth, rather than toxic influent, was the primary problem.
A system audit revealed that the surface aerators were causing excessive evaporative cooling, leading to the slow growth of the nitrifying population. The company acquired a new system that almost completely eliminated aeration-associated evaporative cooling. Within weeks, the nitrifying population returned to normal levels of 2% to 3% of the biomass.
In contrast, MCA of a wastewater treatment plant at an oil refinery revealed a complete loss of the nitrifier population, even after repeated bioaugmentation. Influent streams did not impact ammonia removal in acute toxicity testing, leading to the identification of a chronic toxicity problem caused by a group of compounds accumulating in the biomass.
A microbial community analysis provides a tremendous amount of data — almost too much for routine testing (on a weekly or bi-weekly basis). In situations where operators prefer routine testing to follow key groups, qPCR is an excellent alternative.
Quantitative PCR
qPCR is a selective and sensitive test for estimating the fraction of the biomass made up by specific bacteria of interest and tracking changes in these sub-populations over time. Much in the same way that tracking SVI or sludge depth trends can provide an indicator of future problems, following microbial trends can help head off future challenges.
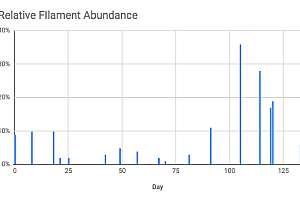
The test is sensitive enough that it can detect low levels of an undesirable filament. If this filament population starts to increase, qPCR can catch the increase before the filament is detected by microscopic examination, lengthening the window of opportunity for corrective action.
Each qPCR test is designed to quantify a particular gene sequence (DNA molecule). If the gene sequence is known, a sensitive and specific test can be developed in a short period of time. Over the last 20 years, qPCR tests have been developed for:
● Ammonia oxidizers
● Nitrite oxidizer
● Nitrate reducers/denitrifiers
● Phosphorus accumulators
● Sulfur oxidizers
● Sulfur reducers
● Many common filaments
● Nocardia foam (Gordonia)
● Antibiotic resistance genes
These tests are flexible and can be customized to fit the needs of an individual plant. qPCR is also a very quick test. When necessary, reports for expedited samples can be returned within 24 hours of receipt. Quick reporting can be instrumental during upsets, particularly when the upset leads to loss of nitrification.
Should the wastewater treatment plant explore the expensive proposition of bioaugmentation with nitrifying cultures? Often, the operators only know that ammonia removal has been reduced or eliminated, but they do not know if the nitrifying population has been killed off or is simply stunned and waiting for conditions to improve. Quick results help operators make informed decisions.
Best practices
For plants looking to add monitoring with molecular diagnostics, the best place to start is by adding a quarterly MCA test, with additional testing during upset conditions. During an upset, it is best to find a laboratory providing a quick turnaround time so that the report is useful. Once a facility understands which key sub-populations impact operations the most, a panel of qPCR tests can be performed weekly or bi-weekly as desired.
About the author: Paul Campbell leads Aster Bio’s molecular biology & biochemistry work. His efforts adopting advanced, high-throughput sequencing, qPCR and other molecular technologies has led to Aster Bio’s Environmental Genomics™ testing platform used to monitor waste treatment system environmental microbial populations and bioprospecting for microbial candidates to include in Aster Bio’s bioaugmentation products. Dr. Campbell has extensive experience with microbial physiology, industrial microbiology and fermentation.